The project EMC2
Monday to Wednesday 07-09/10/2019
Molecular simulation is one of the most dynamic areas of scientific computing. Its field of application is very broad, ranging from theoretical chemistry and drug design to materials science and nanotechnology. Its importance in modern science has been acknowledged by two Nobel Prizes (Kohn & Pople in 1998; Karplus, Levitt & Warshel in 2013). It is also a gold mine of exciting problems for mathematicians and computer scientists.
Molecular simulation can be used as a virtual microscope to study more or less complex molecules with atomic-scale space-time resolution. It can also be used as a tool for computer-aided design (CAD) and the engineering of new molecules, materials and nano-devices.
However, molecular simulation still has important limitations. In particular, the simulation of very large molecular systems, or smaller systems in which electrons interact strongly with each other, remains out of reach today. Overcoming these limitations is extremely difficult. This requires joint breakthroughs in several disciplines, and can, in our opinion, only be achieved through an intensive multidisciplinary effort such as those made possible by ERC-Synergy-type funding.
Our objective is to overcome some of the current limitations in this field and to provide academic communities and industrial companies with new generation, dramatically faster and quantitatively reliable molecular simulation software, to enable those communities to address major technological and societal challenges of the 21st century (in health, energy, and the environment, for example).
Indeed, these challenges require an understanding of the intimate properties of matter at the atomic scale and the development of engineering techniques at that scale. The molecular simulation models, algorithms, and software that we plan to develop in this project will enable the use of CAD (computer-aided design) for new drugs, materials or nano-objects.
An example of progress will be to provide error estimates on the results from numerical simulations (what we call “a posteriori estimators and indicators”). Results can then be supplemented with error bars, which is usual for experimental results. These error bars are crucial for estimating the reliability of the results and their scope.
The high-performance scientific computing component will allow us to integrate new concepts such as algorithms that minimize data transfer between processors, so they can scale on supercomputers, while also reducing energy consumption. These algorithms will be developed and used to accelerate molecular dynamics calculations on future exascale and post-exascale systems (i.e. ten to a thousand times faster than systems currently available). They will also be dedicated to methods from quantum chemistry to efficiently calculate the electronic structures of systems with highly correlated electrons.
This requires dealing with the curse of dimensionality (i.e. the exponential increase in complexity as a function of dimension) inherent in these systems. For this purpose, algorithms and a dedicated computer library using a representation of the data in large dimensions by objects called tensors will be developed, to enable their effective compression, i.e. their representation by simpler objects in small dimensions, while preserving the information. The library will benefit other fields that process large amounts of data, such as artificial intelligence.
This project represents a great opportunity to carry out innovative and cutting-edge research at the interface of chemistry, computer science, and mathematics, which, through major advances in each of these disciplines as well as at their interfaces, will make it possible to discover in silico new molecules and materials.
Ideally, we would like to model an entire cell in all its complexity, paving the way for genomics and personalized medicine at the atomic level. In another area we will develop innovative approaches to simulate new materials such as multilayer 2D materials that have amazing physical properties, many of which are yet to be discovered.
Cliquez ici pour découvrir les participants
Martin AVERSENG
Félix AVIAT
Matthias BEAUPERE
Robert BENDA
Sylvie BOLDO
Eric CANCES
Paul CAZEAUX
Edmond CHEN
Gilles COLRAT
Virginie EHRLACHER
Lucile GARNIER
David GONTIER
Anne Françoise de GUERNY
Nohad GRESH
Laura GRIGORI
Michael HERBST
Charlotte HUBER
Luc-Henri JOLLY
Gaspard KEMLIN
Venera KHOROMSKAYA
Boris KHOROMSKIJ
Daniel KRESSNER
Tony LELIEVRE
Antoine LEVITT
Filippo LIPPARINI
Daniele LOCO
Mitchell LUSKIN
Yvon MADAY
Stéphane MALLAT
Carlo MARCATI
Nastasia MAUGER
Agnieszka MIEDLAR
Pierre MONMARCHE
Matthieu MONTES
Catherine PACHERIE-SIMERAL
Jean-Philip PIQUEMAL
Etienne POLACK
Jay PONDER
Thomas SIMONSON
Isabelle SIMUNIC
Sami SIRAJ-DINE
Benjamin STAMM
Gabriel STOLTZ
Valentin THIRION
Julien TOULOUSE
Martin VOHRALIK
Germain WANG
Jérémy WEISMAN
Agenda
Monday, October 7, 2019
| TIME | EVENT | |
| 09:45 – 10:00 | Registration and welcome breakfast – Registration and welcome breakfast | |
| 10:00 – 11:45 | General presentation of the project – members, objectives, current tasks, synergies between its components | |
| 12:15 – 14:00 | Lunch at the Ardoise on the campus Pierre & Marie Curie (ardoise) | |
| 14:30 – 15:00 | Sylvie Boldo – From mathematics to programs: a verification journey | |
| 15:00 – 15:30 | Benjamin Stamm : Gradient flow finite element discretizations with energy-based adaptivity for the Gross Pitaevskii Equation – We present an effective adaptive procedure for the numerical approximation of the steady-state Gross-Pitaevskii equation which consists of a combination of gradient flow iterations and adaptive finite element mesh refinements. The mesh-refinement is solely based on energy minimization. Numerical tests show that this strategy is able to provide highly accurate results, with optimal convergence rates with respect to the number of freedom. | |
| 15:30 – 16:00 | Coffee break (309, 15-25) | |
| 16:00 – 16:30 | Carlo Marcati – Weighted analytic regularity and exponential convergence for electronic structure calculations | |
| 16:30 – 17:00 | Alston Misquitta – Development of ab initio force fields | |
| 17:00 – 18:00 | Discussions between participants – Discussions between participants |
Tuesday, October 8, 2019
| TIME | EVENT | |
| 09:00 – 09:30 | Jay Ponder – Free energy calculations with AMOEBA | |
| 09:30 – 10:00 | Stéphane Mallat – Wavelet Scattering for Predictions of Molecule Properties – S. Mallat with M. Eickenberg, G. Exarchakis, M. Hirn,Mallat,and L. Thiry We present a machine learning algorithm for the prediction of molecule energies inspired by ideas from density functional theory. Using Gaussian-type orbital functions, we create surrogate electronic densities of the molecule, from which we compute invariant solid harmonic scattering coefficients that account for different types of interactions at different scales. Multi-linear regressions of various physical properties of molecules are computed from these invariant coefficients. Numerical experiments show that these regressions have near state of the art performance, even with relatively few training examples. Predictions over small sets of scattering coe cients can reach a DFT precision while being interpretable. | |
| 10:00 – 10:45 | Coffee break | |
| 10:45 – 11:15 | Venera Khoromskaia – “Tensor numerical methods in quantum chemistry: from Hartree-Fock energies to optical spectra of compact molecules”. | |
| 11:15 – 11:45 | Boris Khoromskij – Range-separated tensor format in bio-molecular modelling and in data scienc | |
| 11:45 – 12:15 | Discussions (309, 15-25) | |
| 12:15 – 14:00 | Lunch Tour Zamansky (24 étage) (salle de réception au 24 eme étage de la tour Zamanski) | |
| 14:00 – 14:30 | Agnieszka Miedlar – Randomized methods and the future of numerical linear algebra | |
| 14:30 – 15:00 | Daniel Kressner : Randomized methods for tensor compression – Probabilistic approaches to low-rank matrix approximation, such as the randomized singular value decomposition, have gained popularity during the last few years, thanks to their simplicity, effectiveness, and flexibility. This talk is concerned with various extensions of these approaches to tensors. In particular, we will focus on the benefits and challenges arising from imposing rank-one structure on the involved random vectors. On the one hand, this can increase the efficiency of randomized methods for tensors, sometimes dramatically. On the other hand, this also significantly complicates the probablistic analysis, e.g., for deriving tail bounds and guaranteeing high accuracy with high probability. This talk is based on joint work with Lana Perisa and Zvonimir Bujanovic. | |
| 15:00 – 15:30 | discussions – discussions | |
| 15:30 – 16:00 | Coffee break | |
| 16:00 – 16:30 | Filippo Lipparini – General linear scaling implementation of polarizable embedding schemes | |
| 16:30 – 17:00 | Martin Vohralik : A posteriori error estimates & adaptivity with balancing error components – We review how to bound the error between the unknown weak solution of a partial differential equation and its computer approximation via a fully computable a posteriori estimate. This allows to certify the numerical simulation result. The particularity of the presented approach is that the derived estimates are valid on each iteration of a linearization procedure, as well as on each algebraic solver iteration; moreover, they allow to distinguish the different error components (discretization/linearization/numerical linear algebra). Fully adaptive algorithms relying on adequate stopping and balancing criteria combined with mesh refinement are designed. Some theoretical results such as efficiency, reliability, robustness with respect to problem or approximation parameters, or convergence and quasi-optimality of the adaptive algorithms are presented and illustrated on numerical experiments. Applications include underground fluid flows and eigenvalue problems. | |
| 17:00 – 18:00 | Discussions between participants | |
| 19:15 – 23:55 | Dinner |
Wednesday, October 9, 2019
| TIME | EVENT | |
| 09:00 – 09:30 | Mitchell Luskin – Momentum Space Methods for the Electronic Structure of 2D Heterostructures | |
| 09:30 – 10:00 | Paul Cazeaux – Twists and misfits: configuration space approaches for relaxation and electronic structure models of 2D heterostructures | |
| 10:00 – 10:45 | Coffee break | |
| 10:45 – 11:15 | Thomas Simonsons : Molecular recognition in complex biomolecular systems – TBA | |
| 11:15 – 11:45 | Julien Toulouse – Strategies for strong correlation in electronic structure theory | |
| 12:30 – 14:30 | Lunch at Tour Zamansky (24th floor) (salle de réception au 24 eme étage de la tour Zamanski) | |
| 14:30 – 18:00 | Discussions between participants – Discussions between participants |
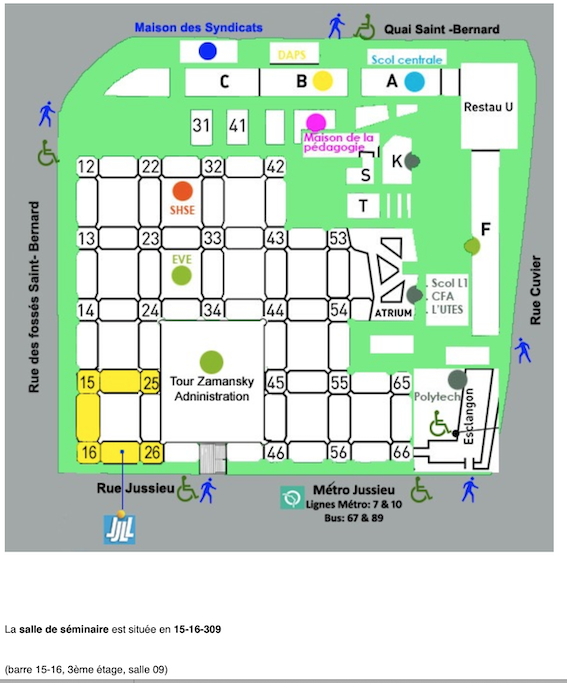
The Kick off meeting took place on the campus Pierre et Marie Curie (Jussieu) of SORBONNE UNIVERSITE
in the Laboratory Jacques-Louis Lions, tower 15-16, 3rd floor, room 309, 4 place Jussieu, 75005 Paris.